 The proposal's effectiveness and efficiency are proved through a variety of experiments. In this paper we thus propose SCOPE, a lightweight privacy preserving approach able to support nodes running the COPE protocol in a secret way, by adopting the Elliptic Curve Cryptography (ECC) homomorphic encryption algorithm. However, a necessary needs of combining data coding mechanism and cryptographic algorithm is raised for both preserving privacy and optimizing bandwidth use at the same time as mentioned above. Hence, while the coding mechanism aims to decrease the bandwidth consumption by aggregating (i.e., coding) multiple packets in the network, the security solutions increase the data quantity. However, the security solutions often exploit the public key schemes which nowadays run more slowly and give the longer encrypted values as the effects of the key size increase to guarantee the algorithm complexity. In order to cope with the mentioned issues, we can apply cryptographic schemes. 
This leads to a serious consequence in leaking the node identity and its private data. The PZC provides important information about metal sorption mechanisms. The COPE data, from the source node through the network to its destination node, can be easily learned by the surrounding nodes, particularly, intersecting nodes and neighbour nodes. This study evaluates different methods to determine points of zero charge (PZCs) on five organic materials, namely maple sawdust, wood ash, peat moss, compost, and brown algae, used for the passive treatment of contaminated neutral drainage effluents. Prot pi provides an accurate tool to calculate isoelectric points of denatured and native proteins that takes charge contributions of multiple subunits, disulfide bonds and posttranslational modifications into account.In this paper, we investigate a simple but effective coding mechanism, namely Coding Opportunistically (COPE), and the privacy violations which are likely to happen to COPE. Figure 2: Correlation of experimental and calculated isoelectric point (pI) of native (blue) and denaturated (green) proteins. The calculated pI values showed reasonably good agreement with experimental ones for all of the six proteins under native and denaturing conditions over a wide range of physico-chemical properties (figure 2). Measured pI were taken from Ui (2006), except: *) Przybylski (2015) and **) Stone-Wolff (1984) Protein 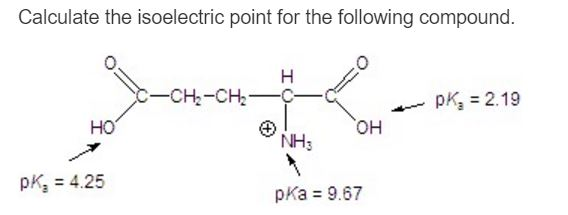
Table 1: Calculated pI compared with measured pI out of literature of native and denaturated proteins. Homo-and heteromultimeric, basic and acidic as well as small and very large proteins were taken to validate the calculated pI (table 1). The calculated and the measured isoelectric point of five proteins with different properties were compared. Figure 1: How to use the algorithm to calculate isoelectric points of native proteins. Choose “native” option in the “Advanced” tab to take advantage of this algorithm (figure 1). An algorithm to determine the isoelectric point of native protein using these average pK values is implemented in Prot pi’s Protein Tool. (2008) summarized 541 pK values of 78 folded proteins measured with NMR which were taken to calculate average pK values for each type of amino acid side chain. On the other hand, amino acid side chain pK values of native proteins where measured using NMR. The isoelectric point corresponds to zero electrokinetic -potential ( 0), i.e. Applications in a variety of fields using.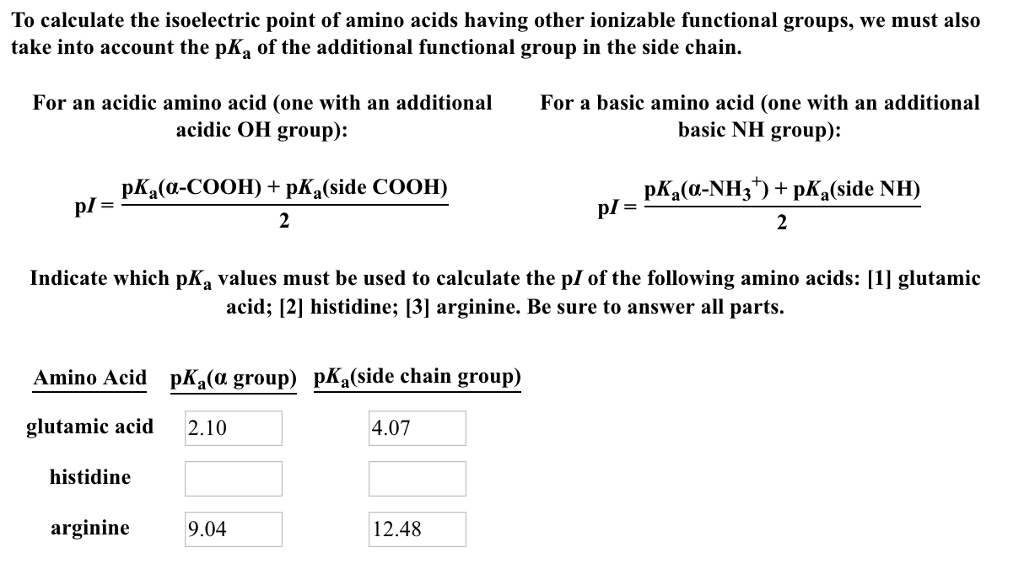 
(1993) published an accurate methode to predict the focusing position of polypeptide in immmobilized pH gradients under denaturing conditions (8 M urea). Derivative methodologies of isoelectric focusing are also discussed including common detection methods used. For denatured proteins, Bjellqvist et al. The usage of appropiate pK values is of crucial importance to caluclate correct pI. For neutral amino acids, the side chains are neutral and the isoelectric point is given simply by the average of the pKa values of carboxylic acid and amine. To determine the pH at which the net charge is zero, the charge on the protein is calculated at multiple pH values using Henderson-Hasselbalch equation, as described by Sillero and Ribeiro (1989). The pI is defined as the pH value at which the positive and negative charges on the protein are balanced and the net charge is zero. The isoelectric point (pI) of native (not denaturated) proteins is important in several separation techniques. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
